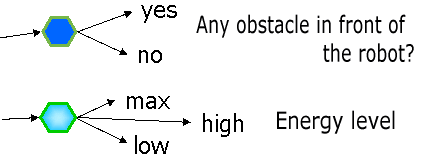
(a) Judgment node
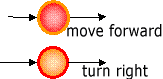
(b) Processing node
An introduction to Genetic Network Programming (GNP) and its coding procedure
<Now revising English sentences>
Evolutionary computation is based on the mechanism of evolution of living
things and applied to many optimization problems, program making etc.
Typical evolutionary computations are:
GA and GP mainly represent solutions using string and tree structures,
respectively.
Genetic Network Programming (GNP)[3,4] has been proposed as a graph-based
evolutionary computation, which is an extension of GA and GP, and also
based on the idea that a graph structure has distinguished abilities to
represent programs because a brain in a living thing is composed of a network
structure.
In this website, the basic strucutre of GNP and how to make program code in C language are introduced.
Template of GNP code ver. 1.0.1 (bug modification 08/07/14)
This is a sample code of GNP which is based on the explanation below.
There are some differences between the sample code and the following explanation,
e.g., variable names, but the functions are the same.
The strucutre of GNP and its features
Component: Node
1. A program of GNP consists of Judgment node (Fig. 1(a))and Processing node(Fig. 2(b)). For example, a judgment node has branch decision functions using sensor inputs, and Processing nodes decises an action to be executed.
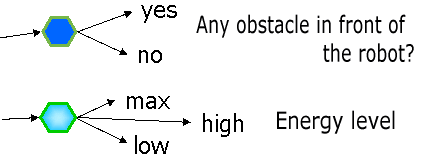 (a) Judgment node |
 (b) Processing node |
| Fig. 1 An example of nodes | |
How to represent node information: Gene strucuture
2. Information of each node is represented by the gene structure (array). *1
| NT | ID | d | CA | CB | CC |
Fig. 2 Gene structure (array)
The gene structure corresponds to the following figure (Fig. 3).
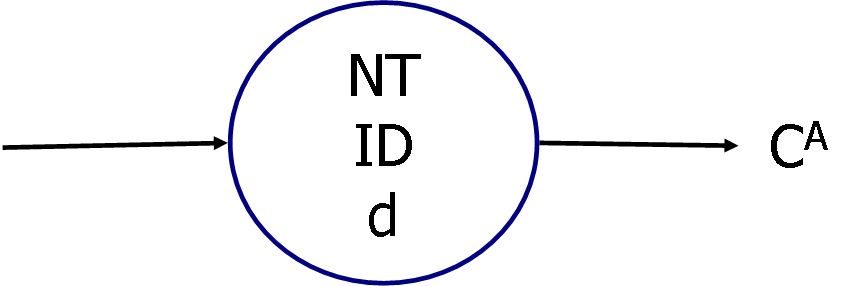
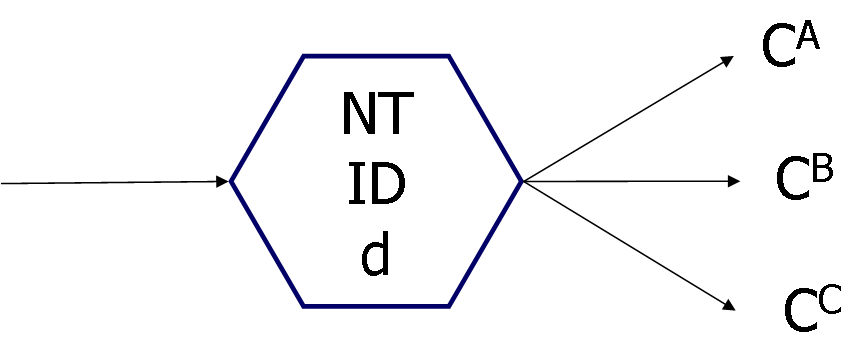

Fig. 3 Correspondence between gene and node
NT:node type (1: Judgment node, 2: Processing node)
ID: node function
In Fig. 1, for example,
NT=1 (Judgment node) and ID=1 shows a judgment node which judges "If
there is any obstacle in front of the robot or not".
NT=1 (Judgment node) and ID=2 shows a judgment node which judges "remaining
energy level".
NT=2 (Processing node) and ID=1 shows a processing node deciding "move
forward" action.
NT=2 (Processing node) and ID=2 shows a processing node deciding "turn
right" action.
d: Time delay needed for node execution, which can be used to consider the
time spend on the judgment/processing. Other function of time delay is
to avoid judgent loop, which is actually very important function.
The setting of time delay depends on a problem. For example,
Time delay d of judgment node = 1
Time delay d of processing node = 5
CA, CB, CC: Node numbers connected from this node. When the total number of nodes
in GNP individual is "N", each node has a unique number from
0 to N-1. Therefore, if CA=15, there is a connection to node number 15.
In the case of judgment node, if the judgment result is "A",
the current node is transferred to node CA. If the judgment result is "B", then next node number is CB. The node judging energy level in Fig. (a), for example, if the energy
is max, next node is CA; if high, next node is CB; if low, the next node is CC.
Processing node has no branch decision function, therefore, after executing
an processing function, e.g., "move forward", "turn right",
etc., the current node is transferred to CA. (CB and CC are not used)
Gene information of one node is represented by the following strucutre
in C language.
struct gene_structure{
int node_type;
int node_func;
int node_delay;
int connect[NUM_BRANCH];
}
/* NUM_BRANCH: max number of branches (connections)*/
*1 In [3,4], there are time delay on node transition, but until now, it has not been used. Therefore, it is omitted in this website.
Directed graph generation
1.Declaration of node variables
GNP program is composed of a large number of judgment nodes and processing
nodes that are connected as a network structure shown in Fig. 3.
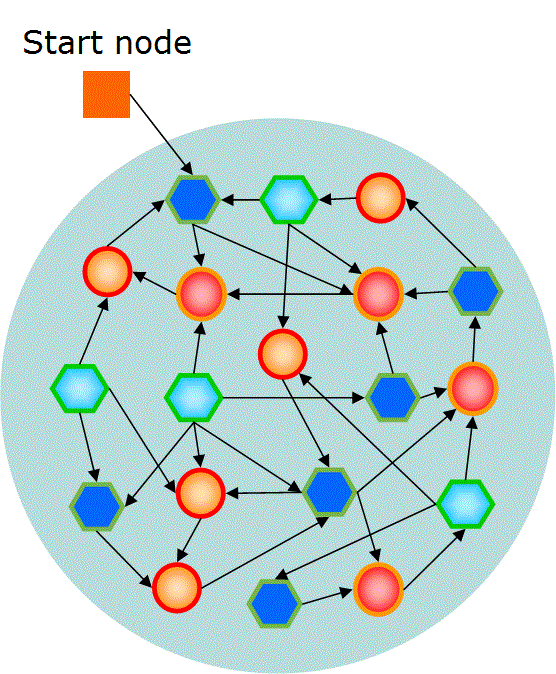
Fig. 3 Basic structure of GNP
First, a variable of a start node information is declared. The start node shows the number of node that is firstly executed in the program.
int initial_node;
Next, we set the number of judgment nodes and processing nodes, which is defined as NODE_SUM. The number of nodes is set depending on the complexity of the problem. For example, if there are four kinds of nodes as Fig. 1, totally 20 nodes are parepared as a first setting, supporsing that five nodes per each kind node function is used. In this case, the gene structure of "NODE_SUM nodes" can be declared by
struct gene_structure gene[NODE_SUM];
Since GNP is an evolutionaty computation, many individuals have to be prepared. Therefore, gene strucutres for all the individuals (the number of individuals is " IND")a are prepared as follows.
| int initial_node[IND]; /* start node numbers for all the individuals */ struct gene_structure gene[IND][NODE_SUM]; /* Judgment nodes and Processing nodes for all the individuals */ |
2. Initialization of the individuals
To create individuals in the first generation, all the variables of nodes are initialized. Node type (judgement/processing) and Node functions are randomly assigned, time delay is set at arbitrary values, and node connections are randomly determined. Suppose that one start node and NODE_SUM nodes including judgment/processing nodes are used, then all the nodes are initialized as follows.
| for (int ind; ind<IND; ind++) { /* Initialization
loop for all the individuals*/ initial_node[ind]=rand()%NODE_SUM; /* Initialization of start (revised 08/07/14) nodes. Randomly assign node numbers that are firstly executed in each indiidual*/ for (int node_num=0; node_num<NODE_SUM; node_num++) { /* Initialization loop for all the nodes */ /* Setting of node type */ if (node_num<JUDGE_SUM) { gene[ind][node_num].node_type=0; /* Prepare the fixed number of judgment nodes *2*/ } else { gene[ind][node_num].node_type=1; /* Prepare the fixed number of processing nodes *2*/ } if (gene[ind][node_num].node_type==0) { gene[ind][node_num].node_func=rand()%JUDGE; /* Judge is the number of node functions. In fig. 1, for example, two functions *3 */ gene[ind][node_num].node_delay=1; /* Time delay of judgment node is one (not limited to one) */ } else { gene[ind][node_num].node_func=rand()%PROCESS; /* PROCESS is the number of node functions. In fig. 1, for example, two functions *3 */ gene[ind][node_num].node_delay=5; /* Time delay of processing node is five(not limited to five)*/ } /* Setting of connection (common to judgment/processing) */ for (int connect_num; connect_num<NUM_BRANCH; connect_num++) { do { gene[ind][node_num].connect[connect_num]=rand()%NODE_SUM; /* Next node number is randomly selected from all the node numbers */ }while (gene[ind][node_num].connect[connect_num]==node_num); /* Self loop is prohibited*/ } } } |
The above program initializes all the nodes and "IND" structures of individuals are created.
*2 Judgment node numbers are from 0 to JUDGE_SUM-1. Processing node numbers are from JUDGE_SUM to NODE_SUM-1.
*3 In the above program, node functions are randomly assigned, but a large
degree of freedom often causes difficulties of search for good solutions.
Therefore, assigning the fixed number of node functions often shows better performance. In Fig. 1, for example, since there are
four kinds of judgment/processing nodes, totally 20 nodes are used preparing
five nodes per each kind. The best number of nodes shoudl be determined
considering the complexity of the problem, and how to find the optimal
number is a remaining problem.
Program execution (node transition)
How to execute individual (number ind (0<=ind<IND, IND: the number of individuals) ) is introduced.
1. A varialbe showing the "current node number" is declared.
int current;
2. Store the first node number according to the start node information.
current=initial_node[ind];
3. Transfer to the next node after executing the function of judgment/processing node.
The program code considering the above is described as follows.
| int current; int judge_result; for (int ind; ind<IND; ind++) { current=initial_node[ind]; for (int step=0; step<TIME_LIMIT;) { if (gene[ind][current].node_type==0) { judge_result=judge(ind, current); /* Execute judgment node and return judgment result judgment function "judge(int ind, int current)" is designed depending on the problem */ } else if (gene[ind][current].node_type==1) { process(ind, current); /* Execute processing function. processing function process(int ind, int current) is designed depending on the problem */ judge_result=0; /* Processing node uses first node branch only */ } step+=gene[ind][current].node_delay; current=gene[ind][current].connect[judge_result]; /* 次のノードへ遷移する */ } } |
Genetic operators
Crossover and mutation are used in GNP.
In the previous research, crossover and mutation are not applied to the
same individuals to avoid destructive changes of the structures, but the
number of individuals created by crossover and those created by mutation
are determined in advance. For example, 120 individuals are created by
crossover and 180 indiiduals are created by mutation.
Crossover
1. Select two individuals (parents) using arbitrary selection method according
to the fitness
2. From node number 1 to N-1, each node is selected as a crossover node
with the probability of Pc.
3. Selected nodes are exchanged between two individuals. For example, node
5 is selected as a crossover node, the gene information of "node 5
in parent1" and that of "node 5 in parent 2" are exchanged.
| for (int node_num=0; node_num<NODE_SUM; node_num++) { /* Each node number
independently executes crossover (uniform crossover) */ if ( ((double)rand()/RAND_MAX)<CROSSOVER_RATE ) { /* determine node number (node_num) is selected or not */ gene_temp[node_num]=gene_parent1[node_num]; /* exchange the genes between two parents */ gene_parent1[node_num]=gene_parent2[node_num]; gene_parent2[node_num]=gene_temp[node_num]; } } |
* The above code of crossover is a little different from the sample program. But, the function is the same.
Mutation
1. Select one individual using arbitrary selection method according to
the fitness
Mutation of node connections:
2. Each branch (connection) is independently selected as a mutation branch
with the probability of Pm.
3. Selected brandh is re-connected to other node.
Mutation of node functions:
4. Each node is independently selected as a mutation node with the probability
of Pm.
5. The functions of selected mutation nodes are randomly changed. However,
processing node is not changed to judgment node, and vice versa.
| for (int node_num=0; node_num<NODE_SUM; node_num++) { /* Mutation is
executed for all the nodes independently */ /* Mutation of connections */ for (int connect_num; connect_num<NUM_BRANCH; connect_num++) {/* Mutation is executed for each branch *4*/ if ( ((double)rand()/RAND_MAX)<MUTATION_RATE ) { /* Determine whether mutation is executed or not according to the mutation rate */ do { gene[ind][node_num].connect[connect_num]=rand()/NODE_SUM; /* next node number is selected from all the node numbers */ }while (gene_temp[mutation][node_num].connect[connect_num]==node_num); /* Prohibit self loop */ } /* Mutation of functions */ if ( ((double)rand()/RAND_MAX)<MUTATION_RATE ) { /* Determine whether mutation is executed or not according to the mutation rate */ if (gene[ind][node_num].node_type==0) { gene[ind][node_num].node_func=rand()/JUDGE; /* If judgment node, judgment function is randomly changed. JUDGE is the number of judgment functions */ } if (gene[ind][node_num].node_type==1) { gene[ind][node_num].node_func=rand()/PROCESS; /* If processing node, processing function is randomly chnges. PROCESS is the number of processing functions */ } } } |
*4 In the above source code, some redundant loop occurs in the mutation of connections of Processing nodes which uses only one connection. The code is described as simple as possible for easy understanding.
References
[1] J. H. Holland: Adaptation in Natural and Artificial Systems, University
of Michigan Press, 1975
[2] J. R. Koza: Genetic Programming: On the Programming of Computers by
Means of Natural Selection, MIT Press. 1992
[3] S. Mabu, K. Hirasawa and J. Hu: A graph-based evolutionary algorithm:
Genetic Network Programming (GNP) and its extension using reinforcement
learning, Evolutionary Computation, Vol. 15, No. 3, pp. 369-398, 2007
[4] 間普 真吾,平澤宏太郎: 遺伝的ネットワークプログラミングのアーキテクチャについて,システム制御情報学会誌,
システム/制御/情報,Vol. 55, No. 11, pp. 480-485, 2011(in Japanese)
Copyright(C) 2014 Shingo Mabu, Graduate School of Science and Enginerring, Yamaguchi University All Rights Reserved.
2014.04.14更新